Note
Go to the end to download the full example code
Benchmark SamplesLoss in 3D
Let’s compare the performances of our losses and backends as the number of samples grows from 100 to 1,000,000.
Setup
import numpy as np
import time
from matplotlib import pyplot as plt
import importlib
import torch
use_cuda = torch.cuda.is_available()
from geomloss import SamplesLoss
MAXTIME = 10 if use_cuda else 0.01 # Max number of seconds before we break the loop
REDTIME = (
2 if use_cuda else 0.01
) # Decrease the number of runs if computations take longer than 2s...
D = 3 # Let's do this in 3D
# Number of samples that we'll loop upon
NS = [
100,
200,
500,
1000,
2000,
5000,
10000,
20000,
50000,
100000,
200000,
500000,
1000000,
]
Synthetic dataset. Feel free to use a Stanford Bunny, or whatever!
def generate_samples(N, device):
"""Create point clouds sampled non-uniformly on a sphere of diameter 1."""
x = torch.randn(N, D, device=device)
x[:, 0] += 1
x = x / (2 * x.norm(dim=1, keepdim=True))
y = torch.randn(N, D, device=device)
y[:, 1] += 2
y = y / (2 * y.norm(dim=1, keepdim=True))
x.requires_grad = True
# Draw random weights:
a = torch.randn(N, device=device)
b = torch.randn(N, device=device)
# And normalize them:
a = a.abs()
b = b.abs()
a = a / a.sum()
b = b / b.sum()
return a, x, b, y
Benchmarking loops.
def benchmark(Loss, dev, N, loops=10):
"""Times a loss computation+gradient on an N-by-N problem."""
# NB: torch does not accept reloading anymore.
# importlib.reload(torch) # In case we had a memory overflow just before...
device = torch.device(dev)
a, x, b, y = generate_samples(N, device)
# We simply benchmark a Loss + gradien wrt. x
code = "L = Loss( a, x, b, y ) ; L.backward()"
Loss.verbose = True
exec(code, locals()) # Warmup run, to compile and load everything
Loss.verbose = False
t_0 = time.perf_counter() # Actual benchmark --------------------
if use_cuda:
torch.cuda.synchronize()
for i in range(loops):
exec(code, locals())
if use_cuda:
torch.cuda.synchronize()
elapsed = time.perf_counter() - t_0 # ---------------------------
print(
"{:3} NxN loss, with N ={:7}: {:3}x{:3.6f}s".format(
loops, N, loops, elapsed / loops
)
)
return elapsed / loops
def bench_config(Loss, dev):
"""Times a loss computation+gradient for an increasing number of samples."""
print("Backend : {}, Device : {} -------------".format(Loss.backend, dev))
times = []
def run_bench():
try:
Nloops = [100, 10, 1]
nloops = Nloops.pop(0)
for n in NS:
elapsed = benchmark(Loss, dev, n, loops=nloops)
times.append(elapsed)
if (nloops * elapsed > MAXTIME) or (
nloops * elapsed > REDTIME and len(Nloops) > 0
):
nloops = Nloops.pop(0)
except IndexError:
print("**\nToo slow !")
try:
run_bench()
except RuntimeError as err:
if str(err)[:4] == "CUDA":
print("**\nMemory overflow !")
else:
# CUDA memory overflows semi-break the internal
# torch state and may cause some strange bugs.
# In this case, best option is simply to re-launch
# the benchmark.
run_bench()
return times + (len(NS) - len(times)) * [np.nan]
def full_bench(loss, *args, **kwargs):
"""Benchmarks the varied backends of a geometric loss function."""
print("Benchmarking : ===============================")
lines = [NS]
backends = ["tensorized", "online", "multiscale"]
for backend in backends:
Loss = SamplesLoss(*args, **kwargs, backend=backend)
lines.append(bench_config(Loss, "cuda" if use_cuda else "cpu"))
benches = np.array(lines).T
# Creates a pyplot figure:
plt.figure()
linestyles = ["o-", "s-", "^-"]
for i, backend in enumerate(backends):
plt.plot(
benches[:, 0],
benches[:, i + 1],
linestyles[i],
linewidth=2,
label='backend="{}"'.format(backend),
)
plt.title('Runtime for SamplesLoss("{}") in dimension {}'.format(Loss.loss, D))
plt.xlabel("Number of samples per measure")
plt.ylabel("Seconds")
plt.yscale("log")
plt.xscale("log")
plt.legend(loc="upper left")
plt.grid(True, which="major", linestyle="-")
plt.grid(True, which="minor", linestyle="dotted")
plt.axis([NS[0], NS[-1], 1e-3, MAXTIME])
plt.tight_layout()
# Save as a .csv to put a nice Tikz figure in the papers:
header = "Npoints " + " ".join(backends)
np.savetxt(
"output/benchmark_" + Loss.loss + "_3D.csv",
benches,
fmt="%-9.5f",
header=header,
comments="",
)
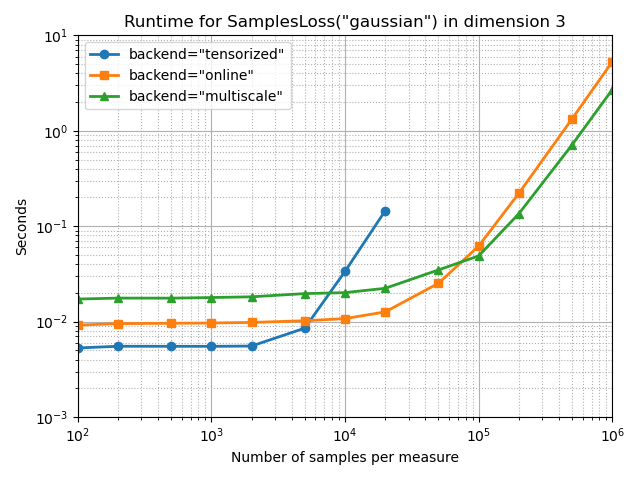
Gaussian MMD, with a small blur
full_bench(SamplesLoss, "gaussian", blur=0.1, truncate=3)
Benchmarking : ===============================
Backend : tensorized, Device : cuda -------------
100 NxN loss, with N = 100: 100x0.005277s
100 NxN loss, with N = 200: 100x0.004129s
100 NxN loss, with N = 500: 100x0.005589s
100 NxN loss, with N = 1000: 100x0.005370s
100 NxN loss, with N = 2000: 100x0.005611s
100 NxN loss, with N = 5000: 100x0.008541s
100 NxN loss, with N = 10000: 100x0.033511s
10 NxN loss, with N = 20000: 10x0.145175s
**
Memory overflow !
Backend : online, Device : cuda -------------
100 NxN loss, with N = 100: 100x0.008343s
100 NxN loss, with N = 200: 100x0.008562s
100 NxN loss, with N = 500: 100x0.008607s
100 NxN loss, with N = 1000: 100x0.008642s
100 NxN loss, with N = 2000: 100x0.008813s
100 NxN loss, with N = 5000: 100x0.009557s
100 NxN loss, with N = 10000: 100x0.010298s
100 NxN loss, with N = 20000: 100x0.012378s
100 NxN loss, with N = 50000: 100x0.024920s
10 NxN loss, with N = 100000: 10x0.063324s
10 NxN loss, with N = 200000: 10x0.224493s
1 NxN loss, with N = 500000: 1x1.332891s
1 NxN loss, with N =1000000: 1x5.293805s
Backend : multiscale, Device : cuda -------------
86x71 clusters, computed at scale = 0.777
100 NxN loss, with N = 100: 100x0.015802s
155x114 clusters, computed at scale = 0.787
100 NxN loss, with N = 200: 100x0.016112s
272x172 clusters, computed at scale = 0.787
100 NxN loss, with N = 500: 100x0.016210s
385x233 clusters, computed at scale = 0.792
100 NxN loss, with N = 1000: 100x0.016580s
493x298 clusters, computed at scale = 0.793
100 NxN loss, with N = 2000: 100x0.016962s
588x392 clusters, computed at scale = 0.794
100 NxN loss, with N = 5000: 100x0.017845s
635x463 clusters, computed at scale = 0.794
100 NxN loss, with N = 10000: 100x0.019896s
673x554 clusters, computed at scale = 0.794
100 NxN loss, with N = 20000: 100x0.021761s
690x626 clusters, computed at scale = 0.794
10 NxN loss, with N = 50000: 10x0.033364s
706x650 clusters, computed at scale = 0.794
10 NxN loss, with N = 100000: 10x0.052962s
708x676 clusters, computed at scale = 0.794
10 NxN loss, with N = 200000: 10x0.144844s
720x696 clusters, computed at scale = 0.794
10 NxN loss, with N = 500000: 10x0.722950s
724x703 clusters, computed at scale = 0.794
1 NxN loss, with N =1000000: 1x2.717151s
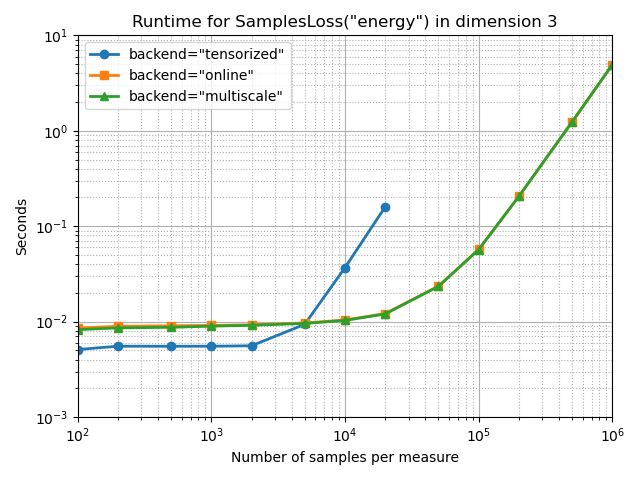
Energy Distance MMD
full_bench(SamplesLoss, "energy")
Benchmarking : ===============================
Backend : tensorized, Device : cuda -------------
100 NxN loss, with N = 100: 100x0.004713s
100 NxN loss, with N = 200: 100x0.005607s
100 NxN loss, with N = 500: 100x0.005603s
100 NxN loss, with N = 1000: 100x0.005613s
100 NxN loss, with N = 2000: 100x0.005653s
100 NxN loss, with N = 5000: 100x0.009382s
100 NxN loss, with N = 10000: 100x0.036823s
10 NxN loss, with N = 20000: 10x0.159420s
**
Memory overflow !
Backend : online, Device : cuda -------------
100 NxN loss, with N = 100: 100x0.007809s
100 NxN loss, with N = 200: 100x0.008116s
100 NxN loss, with N = 500: 100x0.008206s
100 NxN loss, with N = 1000: 100x0.008328s
100 NxN loss, with N = 2000: 100x0.008433s
100 NxN loss, with N = 5000: 100x0.008907s
100 NxN loss, with N = 10000: 100x0.010067s
100 NxN loss, with N = 20000: 100x0.011853s
100 NxN loss, with N = 50000: 100x0.023354s
10 NxN loss, with N = 100000: 10x0.058585s
10 NxN loss, with N = 200000: 10x0.208373s
1 NxN loss, with N = 500000: 1x1.250212s
1 NxN loss, with N =1000000: 1x4.960261s
Backend : multiscale, Device : cuda -------------
100 NxN loss, with N = 100: 100x0.007574s
100 NxN loss, with N = 200: 100x0.008065s
100 NxN loss, with N = 500: 100x0.008037s
100 NxN loss, with N = 1000: 100x0.008175s
100 NxN loss, with N = 2000: 100x0.008347s
100 NxN loss, with N = 5000: 100x0.008614s
100 NxN loss, with N = 10000: 100x0.009972s
100 NxN loss, with N = 20000: 100x0.011715s
100 NxN loss, with N = 50000: 100x0.023245s
10 NxN loss, with N = 100000: 10x0.058860s
10 NxN loss, with N = 200000: 10x0.209811s
1 NxN loss, with N = 500000: 1x1.253326s
1 NxN loss, with N =1000000: 1x4.962973s
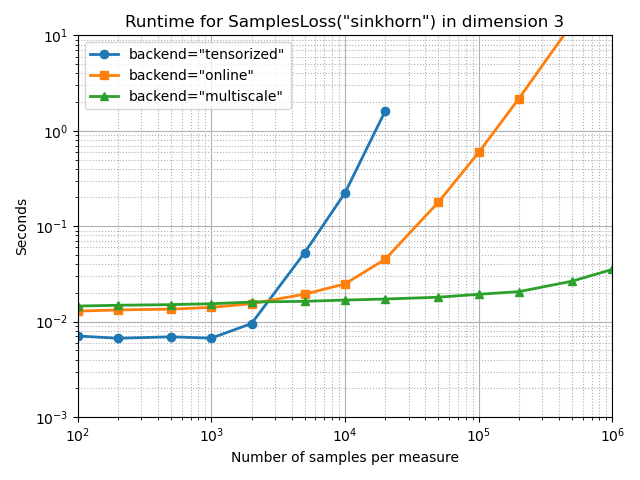
Sinkhorn divergence
With a medium blurring scale, at one twentieth of the configuration’s diameter:
full_bench(SamplesLoss, "sinkhorn", p=2, blur=0.05, diameter=1)
Benchmarking : ===============================
Backend : tensorized, Device : cuda -------------
100 NxN loss, with N = 100: 100x0.006731s
100 NxN loss, with N = 200: 100x0.006892s
100 NxN loss, with N = 500: 100x0.006700s
100 NxN loss, with N = 1000: 100x0.006736s
100 NxN loss, with N = 2000: 100x0.009632s
100 NxN loss, with N = 5000: 100x0.053015s
10 NxN loss, with N = 10000: 10x0.223453s
1 NxN loss, with N = 20000: 1x1.610924s
**
Memory overflow !
Backend : online, Device : cuda -------------
100 NxN loss, with N = 100: 100x0.012718s
100 NxN loss, with N = 200: 100x0.012837s
100 NxN loss, with N = 500: 100x0.013236s
100 NxN loss, with N = 1000: 100x0.013907s
100 NxN loss, with N = 2000: 100x0.015254s
100 NxN loss, with N = 5000: 100x0.019475s
100 NxN loss, with N = 10000: 100x0.024630s
10 NxN loss, with N = 20000: 10x0.045691s
10 NxN loss, with N = 50000: 10x0.179329s
10 NxN loss, with N = 100000: 10x0.595171s
1 NxN loss, with N = 200000: 1x2.181088s
1 NxN loss, with N = 500000: 1x13.347060s
**
Too slow !
Backend : multiscale, Device : cuda -------------
96x86 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.050
Extrapolate from coarse to fine after the last iteration.
100 NxN loss, with N = 100: 100x0.014446s
173x150 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.050
Extrapolate from coarse to fine after the last iteration.
100 NxN loss, with N = 200: 100x0.014587s
393x283 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.050
Extrapolate from coarse to fine after the last iteration.
100 NxN loss, with N = 500: 100x0.014905s
638x432 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.050
Extrapolate from coarse to fine after the last iteration.
100 NxN loss, with N = 1000: 100x0.015152s
945x599 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.050
Extrapolate from coarse to fine after the last iteration.
100 NxN loss, with N = 2000: 100x0.015632s
1344x834 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.050
Extrapolate from coarse to fine after the last iteration.
100 NxN loss, with N = 5000: 100x0.016512s
1632x1061 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.050
Extrapolate from coarse to fine after the last iteration.
100 NxN loss, with N = 10000: 100x0.016505s
1852x1246 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.050
Extrapolate from coarse to fine after the last iteration.
100 NxN loss, with N = 20000: 100x0.016954s
2013x1557 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.050
Extrapolate from coarse to fine after the last iteration.
100 NxN loss, with N = 50000: 100x0.017980s
2070x1768 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.050
Extrapolate from coarse to fine after the last iteration.
100 NxN loss, with N = 100000: 100x0.018911s
2129x1923 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.050
Extrapolate from coarse to fine after the last iteration.
100 NxN loss, with N = 200000: 100x0.020590s
2162x2059 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.050
Extrapolate from coarse to fine after the last iteration.
10 NxN loss, with N = 500000: 10x0.026245s
2186x2118 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.050
Extrapolate from coarse to fine after the last iteration.
10 NxN loss, with N =1000000: 10x0.036538s
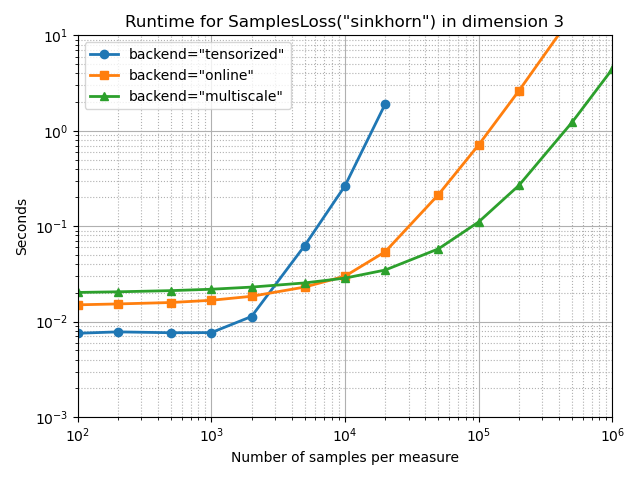
With a small blurring scale, at one hundredth of the configuration’s diameter:
full_bench(SamplesLoss, "sinkhorn", p=2, blur=0.01, diameter=1)
plt.show()
Benchmarking : ===============================
Backend : tensorized, Device : cuda -------------
100 NxN loss, with N = 100: 100x0.007584s
100 NxN loss, with N = 200: 100x0.008110s
100 NxN loss, with N = 500: 100x0.007618s
100 NxN loss, with N = 1000: 100x0.007969s
100 NxN loss, with N = 2000: 100x0.011411s
100 NxN loss, with N = 5000: 100x0.062814s
10 NxN loss, with N = 10000: 10x0.264785s
1 NxN loss, with N = 20000: 1x1.909051s
**
Memory overflow !
Backend : online, Device : cuda -------------
100 NxN loss, with N = 100: 100x0.014850s
100 NxN loss, with N = 200: 100x0.015004s
100 NxN loss, with N = 500: 100x0.015719s
100 NxN loss, with N = 1000: 100x0.016327s
100 NxN loss, with N = 2000: 100x0.018110s
100 NxN loss, with N = 5000: 100x0.023239s
10 NxN loss, with N = 10000: 10x0.030030s
10 NxN loss, with N = 20000: 10x0.053841s
10 NxN loss, with N = 50000: 10x0.214995s
1 NxN loss, with N = 100000: 1x0.715599s
1 NxN loss, with N = 200000: 1x2.631839s
1 NxN loss, with N = 500000: 1x16.137408s
**
Too slow !
Backend : multiscale, Device : cuda -------------
92x81 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.031, 0.016, 0.010
Jump from coarse to fine between indices 5 (σ=0.063) and 6 (σ=0.031).
Keep 1424/7452 = 19.1% of the coarse cost matrix.
Keep 1288/8464 = 15.2% of the coarse cost matrix.
Keep 1677/6561 = 25.6% of the coarse cost matrix.
100 NxN loss, with N = 100: 100x0.020177s
178x159 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.031, 0.016, 0.010
Jump from coarse to fine between indices 5 (σ=0.063) and 6 (σ=0.031).
Keep 3914/28302 = 13.8% of the coarse cost matrix.
Keep 3720/31684 = 11.7% of the coarse cost matrix.
Keep 5085/25281 = 20.1% of the coarse cost matrix.
10 NxN loss, with N = 200: 10x0.020272s
372x295 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.031, 0.016, 0.010
Jump from coarse to fine between indices 5 (σ=0.063) and 6 (σ=0.031).
Keep 14428/109740 = 13.1% of the coarse cost matrix.
Keep 15454/138384 = 11.2% of the coarse cost matrix.
Keep 16963/87025 = 19.5% of the coarse cost matrix.
10 NxN loss, with N = 500: 10x0.020964s
634x441 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.031, 0.016, 0.010
Jump from coarse to fine between indices 5 (σ=0.063) and 6 (σ=0.031).
Keep 34181/279594 = 12.2% of the coarse cost matrix.
Keep 40496/401956 = 10.1% of the coarse cost matrix.
Keep 32045/194481 = 16.5% of the coarse cost matrix.
10 NxN loss, with N = 1000: 10x0.021812s
923x599 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.031, 0.016, 0.010
Jump from coarse to fine between indices 5 (σ=0.063) and 6 (σ=0.031).
Keep 63049/552877 = 11.4% of the coarse cost matrix.
Keep 79857/851929 = 9.4% of the coarse cost matrix.
Keep 52635/358801 = 14.7% of the coarse cost matrix.
10 NxN loss, with N = 2000: 10x0.022992s
1330x842 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.031, 0.016, 0.010
Jump from coarse to fine between indices 5 (σ=0.063) and 6 (σ=0.031).
Keep 112280/1119860 = 10.0% of the coarse cost matrix.
Keep 149232/1768900 = 8.4% of the coarse cost matrix.
Keep 84508/708964 = 11.9% of the coarse cost matrix.
10 NxN loss, with N = 5000: 10x0.025386s
1629x1031 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.031, 0.016, 0.010
Jump from coarse to fine between indices 5 (σ=0.063) and 6 (σ=0.031).
Keep 161430/1679499 = 9.6% of the coarse cost matrix.
Keep 214677/2653641 = 8.1% of the coarse cost matrix.
Keep 116043/1062961 = 10.9% of the coarse cost matrix.
10 NxN loss, with N = 10000: 10x0.029012s
1847x1253 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.031, 0.016, 0.010
Jump from coarse to fine between indices 5 (σ=0.063) and 6 (σ=0.031).
Keep 214621/2314291 = 9.3% of the coarse cost matrix.
Keep 272741/3411409 = 8.0% of the coarse cost matrix.
Keep 157863/1570009 = 10.1% of the coarse cost matrix.
10 NxN loss, with N = 20000: 10x0.036297s
2015x1593 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.031, 0.016, 0.010
Jump from coarse to fine between indices 5 (σ=0.063) and 6 (σ=0.031).
Keep 285047/3209895 = 8.9% of the coarse cost matrix.
Keep 325157/4060225 = 8.0% of the coarse cost matrix.
Keep 230607/2537649 = 9.1% of the coarse cost matrix.
10 NxN loss, with N = 50000: 10x0.058818s
2077x1756 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.031, 0.016, 0.010
Jump from coarse to fine between indices 5 (σ=0.063) and 6 (σ=0.031).
Keep 314777/3647212 = 8.6% of the coarse cost matrix.
Keep 345707/4313929 = 8.0% of the coarse cost matrix.
Keep 273164/3083536 = 8.9% of the coarse cost matrix.
10 NxN loss, with N = 100000: 10x0.110040s
2129x1917 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.031, 0.016, 0.010
Jump from coarse to fine between indices 5 (σ=0.063) and 6 (σ=0.031).
Keep 344704/4081293 = 8.4% of the coarse cost matrix.
Keep 364185/4532641 = 8.0% of the coarse cost matrix.
Keep 323623/3674889 = 8.8% of the coarse cost matrix.
10 NxN loss, with N = 200000: 10x0.265623s
2165x2044 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.031, 0.016, 0.010
Jump from coarse to fine between indices 5 (σ=0.063) and 6 (σ=0.031).
Keep 367386/4425260 = 8.3% of the coarse cost matrix.
Keep 377119/4687225 = 8.0% of the coarse cost matrix.
Keep 369074/4177936 = 8.8% of the coarse cost matrix.
1 NxN loss, with N = 500000: 1x1.221953s
2186x2121 clusters, computed at scale = 0.046
Successive scales : 1.000, 1.000, 0.500, 0.250, 0.125, 0.063, 0.031, 0.016, 0.010
Jump from coarse to fine between indices 5 (σ=0.063) and 6 (σ=0.031).
Keep 381998/4636506 = 8.2% of the coarse cost matrix.
Keep 384808/4778596 = 8.1% of the coarse cost matrix.
Keep 400463/4498641 = 8.9% of the coarse cost matrix.
1 NxN loss, with N =1000000: 1x4.397403s
Total running time of the script: (4 minutes 56.374 seconds)